Note
Go to the end to download the full example code.
MgB2 fatbands
This example shows how to plot the L-projected fatbands of MgB2 using the FATBANDS.nc files produced by abinit with prtdos 3. See also PhysRevLett.86.4656
Open the file (alternatively one can use the shell and abiopen.py FILE -nb to open the file in a jupyter notebook) Note that this file has been produced on a k-path so it’s not suitable for DOS calculations.
import abipy.data as abidata
from abipy import abilab
fbnc_kpath = abilab.abiopen(abidata.ref_file("mgb2_kpath_FATBANDS.nc"))
# To customize the matplotlib marker and its size, use:
fbnc_kpath.marker_spin = {0: "o", 1: "v"}
fbnc_kpath.marker_size = 2
Print file info (dimensions, variables …) Note that prtdos = 3, so LM decomposition is not available.
print(fbnc_kpath)
================================= File Info =================================
Name: mgb2_kpath_FATBANDS.nc
Directory: /home/runner/work/abipy/abipy/abipy/data/refs/mgb2_fatbands
Size: 149.01 kB
Access Time: Sat Jun 6 12:54:27 2026
Modification Time: Sat Jun 6 12:49:13 2026
Change Time: Sat Jun 6 12:49:13 2026
================================= Structure =================================
Full Formula (Mg1 B2)
Reduced Formula: MgB2
abc : 3.086000 3.086000 3.523000
angles: 90.000000 90.000000 120.000000
pbc : True True True
Sites (3)
# SP a b c
--- ---- -------- -------- ---
0 Mg 0 0 0
1 B 0.333333 0.666667 0.5
2 B 0.666667 0.333333 0.5
Abinit Spacegroup: spgid: 191, num_spatial_symmetries: 24, has_timerev: True, symmorphic: False
============================== Electronic Bands ==============================
================================= Structure =================================
Full Formula (Mg1 B2)
Reduced Formula: MgB2
abc : 3.086000 3.086000 3.523000
angles: 90.000000 90.000000 120.000000
pbc : True True True
Sites (3)
# SP a b c
--- ---- -------- -------- ---
0 Mg 0 0 0
1 B 0.333333 0.666667 0.5
2 B 0.666667 0.333333 0.5
Abinit Spacegroup: spgid: 191, num_spatial_symmetries: 24, has_timerev: True, symmorphic: False
Number of electrons: 8.0, Fermi level: 8.700 (eV)
nsppol: 1, nkpt: 78, mband: 8, nspinor: 1, nspden: 1
smearing scheme: none (occopt 1), tsmear_eV: 0.272, tsmear Kelvin: 3157.7
Direct gap:
Energy: 0.595 (eV)
Initial state: spin: 0, kpt: [+0.350, +0.300, +0.000], band: 3, eig: 8.402, occ: 2.000
Final state: spin: 0, kpt: [+0.350, +0.300, +0.000], band: 4, eig: 8.997, occ: 0.000
Fundamental gap:
Energy: 0.058 (eV)
Initial state: spin: 0, kpt: K [+0.333, +0.333, +0.000], band: 4, eig: 8.700, occ: 0.000
Final state: spin: 0, kpt: [+0.147, +0.000, +0.000], band: 4, eig: 8.758, occ: 0.000
Bandwidth: 14.314 (eV)
Valence maximum located at kpt index 27:
spin: 0, kpt: K [+0.333, +0.333, +0.000], band: 4, eig: 8.700, occ: 0.000
Conduction minimum located at kpt index 5:
spin: 0, kpt: [+0.147, +0.000, +0.000], band: 4, eig: 8.758, occ: 0.000
TIP: Use `--verbose` to print k-point coordinates with more digits
=============================== Fatbands Info ===============================
prtdos: 3, prtdosm: 0, mbesslang: 5, pawprtdos: 0, usepaw: 0
nsppol: 1, nkpt: 78, mband: 8
Idx Symbol Reduced_Coords Lmax Ratsph [Bohr] Has_Atom
----- -------- ----------------------- ------ --------------- ----------
0 Mg 0.00000 0.00000 0.00000 4 2 Yes
1 B 0.33333 0.66667 0.50000 4 2 Yes
2 B 0.66667 0.33333 0.50000 4 2 Yes
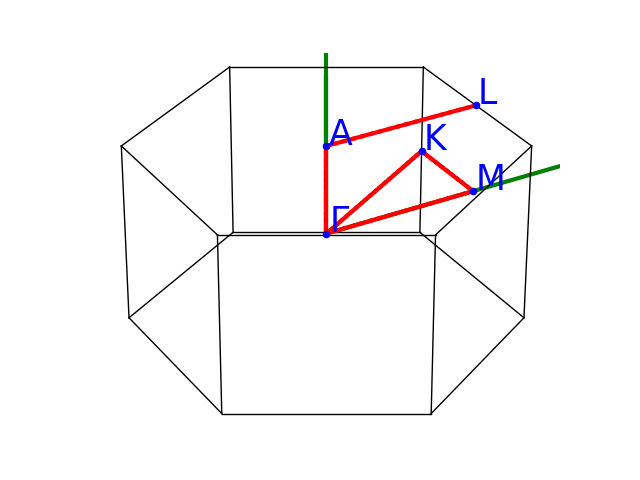
Plot the k-points belonging to the path. fbnc_kpath.ebands.kpoints.plotly()
NC files have contributions up to L=4 (g channel) but here we are intererested in s,p,d terms only so we use the optional argument lmax
lmax = 2
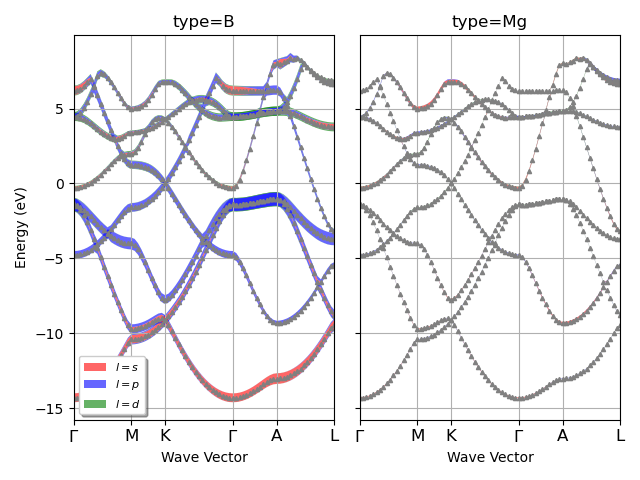
Plot the electronic fatbands grouped by atomic type. Can use l_list to select only particular l-values
fbnc_kpath.plot_fatbands_typeview(lmax=lmax, tight_layout=True, l_list=None)
For the plotly version use:
# fbnc_kpath.plotly_fatbands_typeview(lmax=lmax)
Plot the electronic fatbands grouped by L.
# fbnc_kpath.plot_fatbands_lview(lmax=lmax, tight_layout=True)
For the plotly version use:
# fbnc_kpath.plotly_fatbands_lview(lmax=lmax)
Now we read another FATBANDS file produced on a 18x18x18 k-mesh This file can be used to plot the projected-DOS
fbnc_kmesh = abilab.abiopen(abidata.ref_file("mgb2_kmesh181818_FATBANDS.nc"))
Let’s print the object
print(fbnc_kmesh)
# fbnc_kmesh.ebands.kpoints.plot()
================================= File Info =================================
Name: mgb2_kmesh181818_FATBANDS.nc
Directory: /home/runner/work/abipy/abipy/abipy/data/refs/mgb2_fatbands
Size: 480.70 kB
Access Time: Sat Jun 6 12:54:27 2026
Modification Time: Sat Jun 6 12:49:13 2026
Change Time: Sat Jun 6 12:49:13 2026
================================= Structure =================================
Full Formula (Mg1 B2)
Reduced Formula: MgB2
abc : 3.086000 3.086000 3.523000
angles: 90.000000 90.000000 120.000000
pbc : True True True
Sites (3)
# SP a b c
--- ---- -------- -------- ---
0 Mg 0 0 0
1 B 0.333333 0.666667 0.5
2 B 0.666667 0.333333 0.5
Abinit Spacegroup: spgid: 191, num_spatial_symmetries: 24, has_timerev: True, symmorphic: False
============================== Electronic Bands ==============================
================================= Structure =================================
Full Formula (Mg1 B2)
Reduced Formula: MgB2
abc : 3.086000 3.086000 3.523000
angles: 90.000000 90.000000 120.000000
pbc : True True True
Sites (3)
# SP a b c
--- ---- -------- -------- ---
0 Mg 0 0 0
1 B 0.333333 0.666667 0.5
2 B 0.666667 0.333333 0.5
Abinit Spacegroup: spgid: 191, num_spatial_symmetries: 24, has_timerev: True, symmorphic: False
Number of electrons: 8.0, Fermi level: 6.851 (eV)
nsppol: 1, nkpt: 370, mband: 8, nspinor: 1, nspden: 1
smearing scheme: cold smearing of N. Marzari with minimization of the bump (occopt 4), tsmear_eV: 0.816, tsmear Kelvin: 9473.2
=============================== Fatbands Info ===============================
prtdos: 3, prtdosm: 0, mbesslang: 5, pawprtdos: 0, usepaw: 0
nsppol: 1, nkpt: 370, mband: 8
Idx Symbol Reduced_Coords Lmax Ratsph [Bohr] Has_Atom
----- -------- ----------------------- ------ --------------- ----------
0 Mg 0.00000 0.00000 0.00000 4 2 Yes
1 B 0.33333 0.66667 0.50000 4 2 Yes
2 B 0.66667 0.33333 0.50000 4 2 Yes
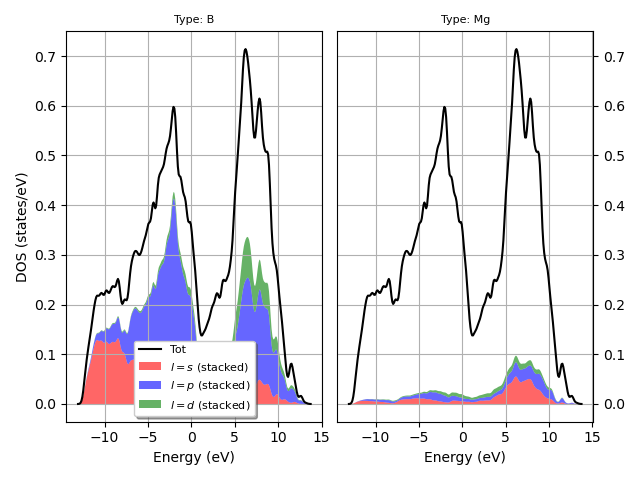
Plot the L-PJDOS grouped by atomic type.
fbnc_kmesh.plot_pjdos_typeview(lmax=lmax, tight_layout=True)
For the plotly version use:
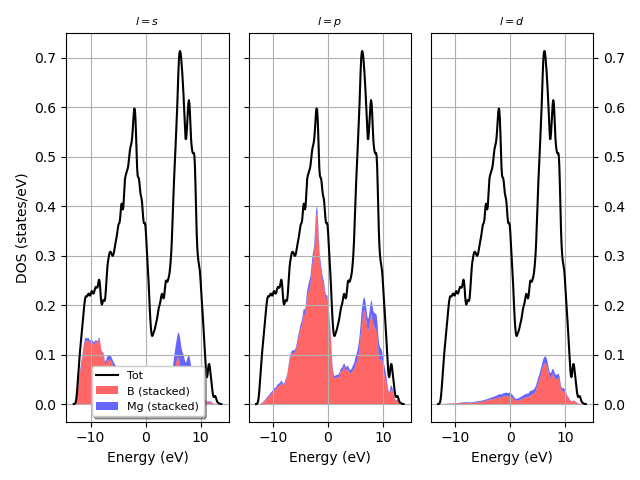
Plot the L-PJDOS grouped by L.
fbnc_kmesh.plot_pjdos_lview(lmax=lmax, tight_layout=True)
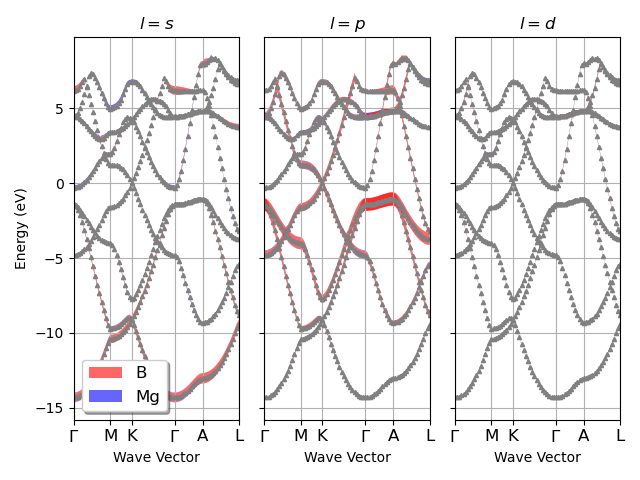
For the plotly version use:
Now we use the two netcdf files to produce plots with fatbands + PJDOSEs. The data for the DOS is taken from pjdosfile. sphinx_gallery_thumbnail_number = 6
fbnc_kpath.plot_fatbands_with_pjdos(pjdosfile=fbnc_kmesh, lmax=lmax, view="type", tight_layout=True)
For the plotly version use:
fbnc_kpath.plotly_fatbands_with_pjdos(pjdosfile=fbnc_kmesh, lmax=lmax, view="type")
fatbands + PJDOS grouped by L
fbnc_kpath.plot_fatbands_with_pjdos(pjdosfile=fbnc_kmesh, lmax=lmax, view="lview", tight_layout=True)
For the plotly version use:
fbnc_kpath.plotly_fatbands_with_pjdos(pjdosfile=fbnc_kmesh, lmax=lmax, view="lview")
# fbnc_kpath.close()
# fbnc_kmesh.close()
Total running time of the script: (0 minutes 2.663 seconds)